Information about the Theoretical and Computational Biophysics Group.
 The Theoretical and Computational Biophysics Group (TCBG) research focuses on the structure and function of supramolecular systems in the living cell, as well as on the development of new algorithms and efficient computing tools for physical biology. We bring the most advanced molecular modeling, bioinformatics, and computational technologies to bear on questions of biomedical relevance. We extend, refine, and deliver these technologies in response to experimental progress and the emerging needs of the wide biomedical research community. We magnify the impact of our work through direct collaboration with experimental researchers, the distribution of cutting-edge and user-friendly software, and extensive training, service, and dissemination efforts.
The Theoretical and Computational Biophysics Group (TCBG) research focuses on the structure and function of supramolecular systems in the living cell, as well as on the development of new algorithms and efficient computing tools for physical biology. We bring the most advanced molecular modeling, bioinformatics, and computational technologies to bear on questions of biomedical relevance. We extend, refine, and deliver these technologies in response to experimental progress and the emerging needs of the wide biomedical research community. We magnify the impact of our work through direct collaboration with experimental researchers, the distribution of cutting-edge and user-friendly software, and extensive training, service, and dissemination efforts.
Our investigations - in collaboration with experimental laboratories in universities, research institutions and industry across the U.S. and around the world - explore the physical mechanisms underlying the transformation of light energy into electrical membrane potentials and the synthesis of ATP in photosynthetic systems, as well as the storage and control of genetic information in living cells. The TCBG develops a theory of the classical and quantum dynamical motion of biopolymers which utilizes numerical experiments, non-equilibrium statistical mechanics, elasticity theory, and the theory of disordered systems. For more information go here.
We have produced ground-breaking insights into biomolecular processes coupled with mechanical force, bioelectronic processes in metabolism and vision, and with the function and mechanism of membrane proteins. We are committed and work towards further advancement of:
- Molecular modeling tools which can integrate structural information with bioinformatics databases and molecular dynamics simulations, and which can be used by a wide audience;
- High-performance molecular visualization and simulation software, capable of modeling biomolecules in realistic environments of 100,000,000 atoms or more;
- Conceptual and methodological foundations of molecular modeling in the fields of quantum biology, mechanobiology, and interactive modeling;
- Biomedical science through collaborations between theoretical and experimental researchers;
- Support of the entire research process and training through a web-enabled collaborative environment; and
- Service, training, and dissemination by leveraging web-based molecular graphics and integrated modeling technologies.
Our Principal Investigator is Emad Tajkhorshid, J. W. Hastings Endowed Chair of Biochemistry and Director of NIH Center for Macromolecular Modeling and Bioinformatics.
Click the tabs below or check out our website here.
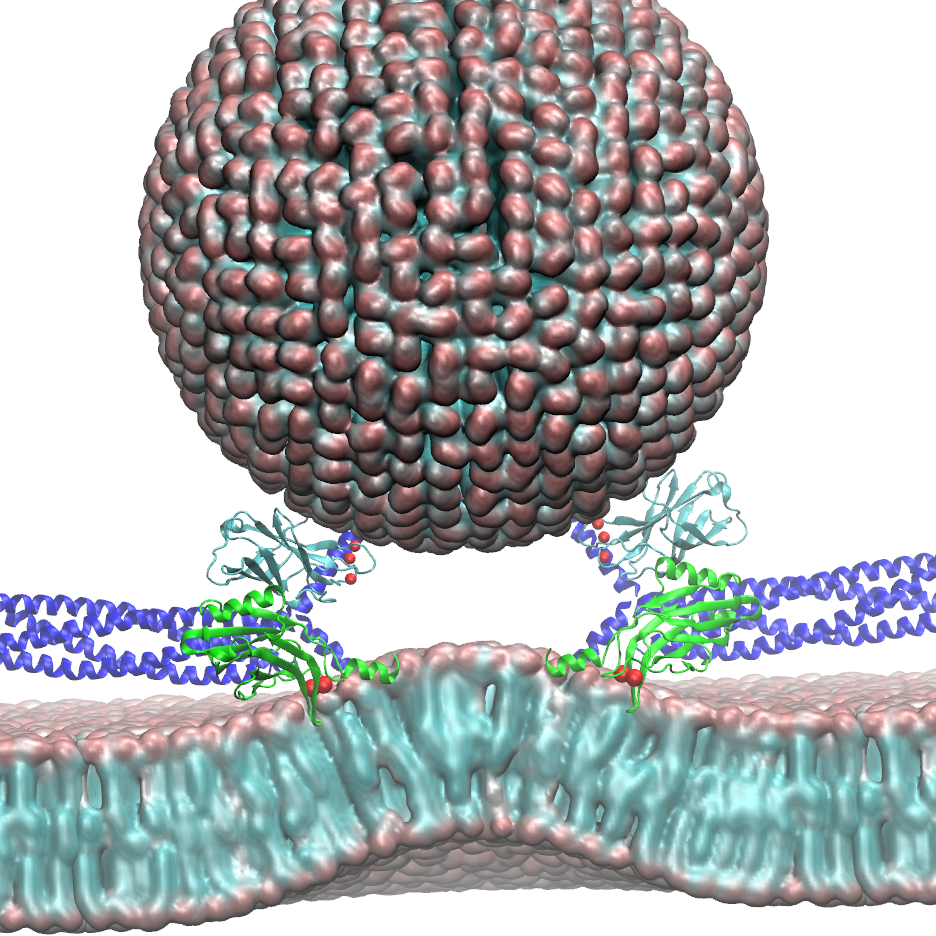 The brain is the source of thoughts, perceptions, emotions, memories, and actions. Neural signaling, the foundation of brain activity, must be precisely regulated to prevent neuronal disorders that may cause Parkinson's disease, schizophrenia, compulsive behaviors, and addiction. Such a precise regulation is achieved by key signaling proteins, voltage-gated sodium and potassium channels for electrical signaling and calcium - bound synaptotagmin for chemical signaling. Here, innovations in computer simulation techniques will be used to investigate the molecular mechanism of neural firing induced by voltage-gated sodium and potassium channels and membrane fusion triggered by synaptotagmin.
The brain is the source of thoughts, perceptions, emotions, memories, and actions. Neural signaling, the foundation of brain activity, must be precisely regulated to prevent neuronal disorders that may cause Parkinson's disease, schizophrenia, compulsive behaviors, and addiction. Such a precise regulation is achieved by key signaling proteins, voltage-gated sodium and potassium channels for electrical signaling and calcium - bound synaptotagmin for chemical signaling. Here, innovations in computer simulation techniques will be used to investigate the molecular mechanism of neural firing induced by voltage-gated sodium and potassium channels and membrane fusion triggered by synaptotagmin.