Information about the Bell Lab.

Research in the Bell lab is focused on understanding why individual animals behave differently from each other. Even an individual fish, for example, behaves differently from other fish, through time and across situations. We study the proximate and ultimate causes of individual variation in the three-spined stickleback. Work in the Bell lab is focused on understanding behavioral variation both from a proximate and ultimate perspective. To that end, we study the neuroendocrine and molecular mechanisms that contribute to behavior, and the selective pressures that maintain variation within natural populations.
The Bell lab is involved in a collaborative project in our theme at the Carl Woese Institute for Genomic Biology to test the hypothesis that there is a ‘genetic toolkit’ for social behavior. We are comparing neurogenomic responses to social opportunities and challenges among honeybees, sticklebacks, and mice (PIs: Gene Robinson and Lisa Stubbs).
Our Principal Investigator is Alison Marie Bell.
Check out at our website here.
 Over the past several years the lab has grown increasingly interested in the possibility that the environment experienced by parents can influence the phenotypic development of their offspring (transgenerational plasticity). We have shown that female sticklebacks are an important source of variation in their offspring, not just via their genes, but via the environment, they provide for their offspring. In particular, we found that mothers exposed to threatening stimuli such as a predator produce offspring with altered behavior (Giesing et al 2011), stress response (Giesing et al 2011) and learning (Roche et al 2012). When testing the hypothesis that predator-mediated maternal effects are mediated through the glucocorticoid stress response system,
Over the past several years the lab has grown increasingly interested in the possibility that the environment experienced by parents can influence the phenotypic development of their offspring (transgenerational plasticity). We have shown that female sticklebacks are an important source of variation in their offspring, not just via their genes, but via the environment, they provide for their offspring. In particular, we found that mothers exposed to threatening stimuli such as a predator produce offspring with altered behavior (Giesing et al 2011), stress response (Giesing et al 2011) and learning (Roche et al 2012). When testing the hypothesis that predator-mediated maternal effects are mediated through the glucocorticoid stress response system,  Sticklebacks are particularly interesting subjects for studying transgenerational plasticity mediated by parental behavior because fathers provide care for their developing offspring, and we’ve been interested in the possibility that the early environmental provided by fathers is important for offspring development. When she was a Ph.D. student in the lab,
Sticklebacks are particularly interesting subjects for studying transgenerational plasticity mediated by parental behavior because fathers provide care for their developing offspring, and we’ve been interested in the possibility that the early environmental provided by fathers is important for offspring development. When she was a Ph.D. student in the lab,  Parental care is fascinating for countless reasons – it’s critical for fitness, there is tremendous variation in the specific forms and manifestations of parental care across organisms, and it’s a highly dynamic interaction between relatives. One of the things that make sticklebacks – like many other fishes – interesting is that they exhibit paternal care, where the father is solely responsible for the care of the offspring. We are very interested in the evolution, inheritance, neurogenomics, and plasticity of paternal care in sticklebacks.
Parental care is fascinating for countless reasons – it’s critical for fitness, there is tremendous variation in the specific forms and manifestations of parental care across organisms, and it’s a highly dynamic interaction between relatives. One of the things that make sticklebacks – like many other fishes – interesting is that they exhibit paternal care, where the father is solely responsible for the care of the offspring. We are very interested in the evolution, inheritance, neurogenomics, and plasticity of paternal care in sticklebacks. We’ve found that some male sticklebacks provide more care than others (Stein 2012, Stein and Bell 2015), the quantity of care received influences offspring traits (McGhee and Bell 2014), and that paternal care might have contributed to the stickleback radiation (Stein and Bell 2019). In a recent study, we found that individual differences in a key parental behavior, fanning, is highly heritable among individual male sticklebacks from the same population (Bell et al 2018). These results suggest that variation in paternal care is tractable for further genetic dissection, e.g. via QTL or GWAS.
We’ve found that some male sticklebacks provide more care than others (Stein 2012, Stein and Bell 2015), the quantity of care received influences offspring traits (McGhee and Bell 2014), and that paternal care might have contributed to the stickleback radiation (Stein and Bell 2019). In a recent study, we found that individual differences in a key parental behavior, fanning, is highly heritable among individual male sticklebacks from the same population (Bell et al 2018). These results suggest that variation in paternal care is tractable for further genetic dissection, e.g. via QTL or GWAS.  Behavioral traits offer exciting new opportunities for integrating quantitative genetics with results from studies of molecular mechanisms and genomics (Bell and Dochtermann 2015). The rationale is that in order to fully understand behavioral traits, they require consideration of both variation and plasticity, and the fixed and dynamic sides of the genome. The growing field of animal personality research prompts this integration because it has brought questions about how and why variation and plasticity coexist to the forefront of thinking in evolutionary and behavioral ecology. We would argue, moreover, that these features of behavioral traits (variation among individuals and plasticity within individuals) are shared with other fitness-related traits of interest to biologists, including health-related and physiological traits, therefore our findings have broad relevance (Bengston et al 2018).
Behavioral traits offer exciting new opportunities for integrating quantitative genetics with results from studies of molecular mechanisms and genomics (Bell and Dochtermann 2015). The rationale is that in order to fully understand behavioral traits, they require consideration of both variation and plasticity, and the fixed and dynamic sides of the genome. The growing field of animal personality research prompts this integration because it has brought questions about how and why variation and plasticity coexist to the forefront of thinking in evolutionary and behavioral ecology. We would argue, moreover, that these features of behavioral traits (variation among individuals and plasticity within individuals) are shared with other fitness-related traits of interest to biologists, including health-related and physiological traits, therefore our findings have broad relevance (Bengston et al 2018). When our lab was established, the stickleback genome had just been sequenced and we saw opportunities to use emerging genomic technologies to uncover the proximate causes of individual differences in behavior. A genome-wide approach was appealing because the behaviors of interest are influenced by many interacting genes of small effect and candidate genes for the behaviors explained only a small fraction of the total variation. Moreover, measuring gene expression (rather than quantifying static DNA sequence variation) was attractive because while there is a heritable component to behavioral traits, they are also plastic, and change with the environment. Instead of focusing on the static side of the genome (DNA sequence), initial studies in the lab probed behavioral variation at the molecular level by measuring brain genome-wide expression in order to learn how the genome dynamically responds to the environment. We found that the stickleback genome is highly responsive to many important aspects of the environment such as predation risk (Sanogo et al 2011), a territorial intrusion (Bukhari et al 2017) and a courtship opportunity (Sanogo and Bell 2016). An important element of this line of work includes tool development for this system.
When our lab was established, the stickleback genome had just been sequenced and we saw opportunities to use emerging genomic technologies to uncover the proximate causes of individual differences in behavior. A genome-wide approach was appealing because the behaviors of interest are influenced by many interacting genes of small effect and candidate genes for the behaviors explained only a small fraction of the total variation. Moreover, measuring gene expression (rather than quantifying static DNA sequence variation) was attractive because while there is a heritable component to behavioral traits, they are also plastic, and change with the environment. Instead of focusing on the static side of the genome (DNA sequence), initial studies in the lab probed behavioral variation at the molecular level by measuring brain genome-wide expression in order to learn how the genome dynamically responds to the environment. We found that the stickleback genome is highly responsive to many important aspects of the environment such as predation risk (Sanogo et al 2011), a territorial intrusion (Bukhari et al 2017) and a courtship opportunity (Sanogo and Bell 2016). An important element of this line of work includes tool development for this system.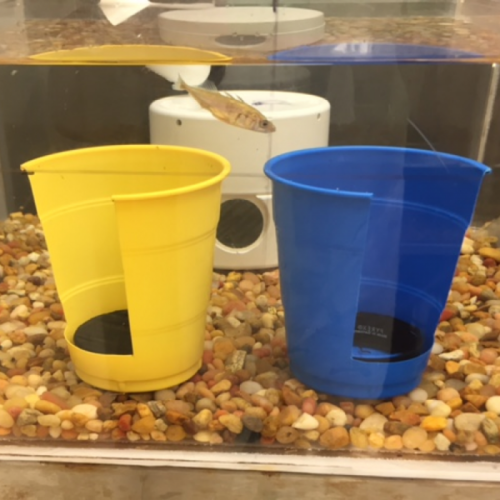  A theme running throughout all of the work in the lab has to do with individual variation. Almost 20 years ago, I had the privilege of working with
A theme running throughout all of the work in the lab has to do with individual variation. Almost 20 years ago, I had the privilege of working with